In 1997, Jill Banfield was a young geologist at the University of Wisconsin–Madison who had become intensely interested in microbiology. Together with Ken Nealson, a microbiologist who had become interested in geology, she organized a groundbreaking geomicrobiology conference in Alta, Utah, inviting the small group of experts whose work in the respective fields overlapped.
Bringing the two camps together wasn’t an easy—or obvious—thing to do. But Banfield had become convinced that you couldn’t study the Earth without also studying the microscopic creatures that were making and remaking it.
Things did not get off to an auspicious start.
“We get there, and it’s this run-down, almost-trailer-park ski lodge,” Banfield recalls. “And I walked in, and the room that they set aside for us to have this meeting in had floor-to-ceiling glass windows, with no curtains, and we had huge slide projectors. With a huge crack across the window. So, I called up this postdoc, and I said, ‘Go out to the hardware store and buy duct tape and black plastic and bring it on the plane with you.’” Then she noticed that the roof was leaking. The snowmelt was dripping where the speakers would be standing. There was no hot water. And, there was no booze.
“This is a disaster,” Banfield remembers thinking, “and everybody is grumbling, and it’s going to be a complete and utter train wreck, and my life is over, my career is over … ”
But someone drove out to fetch beer, and by morning, everyone was laughing. Ultimately, the less-than-idyllic setting helped bond the conferees, who would, together, advance the little-known discipline. Talks from Alta became chapters in a founding text on the subject, Geomicrobiology: Interactions Between Microbes and Minerals.
Helping to forge the modern field of geomicrobiology was just the start. Banfield has since expanded her scope of inquiry dramatically, from mineral weathering to uncovering new microbe species that change how we think about life itself; coming up with ways to use microbes to clean up pollutants, make more fertile soil, and capture carbon; and organizing the first meetings on CRISPR. Most recently, she and her colleagues at Berkeley’s Innovative Genomics Institute have become fascinated by what they’ve dubbed “Borgs”—mysterious pieces of DNA inside of microbes deep in California mud, and which might play a crucial role in allaying global warming.
Banfield is highly energetic and focused, relentlessly excited about data. On a typical day (pre-COVID-19), she walks to work from her home in Berkeley. She keeps her schedule clear before 10 a.m., saving early-morning time for uninterrupted work. Much of the 10–5 is occupied by meetings, data analysis, and exploring new project ideas.
But if you can stick around after 5 p.m., you can have her undivided, caffeine-augmented attention for hours. Warning: She may forget that, unlike her, you need to eat.
“Her voice is strong in my head because it was so many hours we spent working together, side-by-side, drinking green tea, me contemplating if she was an autotroph or how she possibly got enough nutrients to sustain herself,” says a former member of Berkeley’s Banfield Laboratory who admits to sneaking off to nosh a granola bar during their chats.
Many people I talked to for this story commented on Banfield’s intense focus, expressed through a piercing, glacier-blue gaze. I can corroborate: I first met Banfield in 2020 to talk about a Nature paper she co-authored on huge phages. Under her unsparing eye contact, I felt a bit like a chicken under a rotisserie lamp. But then she described her findings—enormous bacteria-infecting viruses, everywhere—with a kind of lucidity that brought them to sparkling life. I emerged from the meeting giddy, briefly wishing I’d gotten my doctorate in virology, instead of genetics.
Later, I was excited when I got the chance to work with her on a written proposal. Her first draft was poetic and looping, the research plan building in a spiraling manner, the concluding sentence hitting like a plot twist. Scientists typically form research plans by starting at A and then moving to B, then C, in a logical half-step progression. Banfield lands near Z as if it comes to her all at once, and then goes back, filling in and making explicit the logic and progression for the rest of us.
Despite my fangirling, I put off interviewing Banfield for this profile, talking to her colleagues and mentees first, in the hopes of learning enough about her work to avoid asking a Bad Question. And according to those who know her well, Banfield believes there is such a thing. Beginner questions aren’t bad. In fact, as a self-taught microbiologist, Banfield spent two years studying microbiology textbooks—“I had no idea there was sugar in DNA!” she recalls happily—and peppering her colleagues with beginner questions.
Bad Questions are ones that reveal a lack of thought or attention, and Banfield is not known for suffering fools or mincing words. “She’ll tell you what she thinks,” says a colleague. “Not meanly, but very direct.” When there’s a disagreement about data, she pulls no punches: “She wants you to make your argument,” says another colleague. “She’s curious. It’s not personal. She fights so that she can learn something.”
It was as a young girl growing up in New South Wales, Australia, that Banfield first became fascinated by geology. On walks, she saw all kinds of rocks and began to understand that they could tell stories: Rocks could help you understand the world and how it has changed over time. She was especially interested in minerals: “I was captivated by their colors, but as soon as I started to learn about them, I wanted to understand how they were put together, how the atoms were arranged.”
Banfield did her undergraduate studies at the Australian National University (ANU) under the mentorship of Tony Eggleton.
“I would give a lecture, and after the lecture, she would come up and say, ‘I didn’t quite get such and such, could you go over it again?’” Eggleton says. “And after a couple seconds, a sudden glaze would come over her face.” Perhaps she was not really interested, and had just asked a question to be polite, Eggleton thought. Then he realized: “It only took a few words, and she got it.”
Between her bachelor’s and master’s degrees at ANU, Banfield spent a brief period working at a mining company. “They were going to send me off to Western Australia to look in gold mines and towns in the Outback, and I was like, ‘No, this is not what my life is meant to be,’” Banfield says. “Sometimes the thing that teaches you what you want to do with your life is the experience of finding out what you don’t want to do.”
Working with Eggleton, Banfield learned techniques to visualize the atomic structures of minerals. “Her real skill initially was electron microscopy,” says Nealson. “She could see things nobody else could see—and a lot of them just didn’t fit.”
Banfield realized that the kinds of things she was seeing in minerals couldn’t be explained by standard, inanimate chemical reactions. They were the chemical fingerprints of living things.

She began to study the weathering of granite and the minerals that comprise it. “She found a little piece of mica, so she split it open, put it in the scanning electron microscope, and had a look at it,” Eggleton recalls. “She found some funny little things shaped like donuts, a couple of microns across. No one knew what they were.” They wondered if the donuts were actually fossilized bacteria, weathering the mica. “That’s where she realized that this was something nobody knew was going on inside rocks.… That was the beginning of her microbiology.”
In 1986, with the support of a Fulbright fellowship, Banfield moved to the United States to do her graduate work in geology at Johns Hopkins University, studying how rocks transform under heat and pressure. In 1990, she joined the geology department at the University of Wisconsin–Madison as an assistant professor. During her 11 years there, she studied the intricacies of mineral weathering, crystal growth, mineral and rock transformation, and how microbes shape these processes.
Though Banfield’s work was published in prestigious journals, the microbiology field was skeptical of her work. She had been trained as a geologist, after all, so her unexpected findings were assumed to be artifacts of poor technique. “Almost none of the people of the microbiologists at Madison were doing the work that she was doing, and they didn’t understand it,” says Nealson. “They didn’t want to understand it, in a sense. They didn’t think it was important.”
Even after she received a 1999 MacArthur “genius grant,” many remained dubious. They tried to prove her wrong, but couldn’t, says Nealson. “This happens to Jill over and over.”
Banfield’s biggest innovation to date—the technique known as genome-resolved metagenomics—came after her move to Berkeley in 2001. Metagenomics is the evolution of new ways of studying microbial communities pioneered in the 1980s by microbiologist Norman Pace. At that time, almost everything we knew about microorganisms came from growing a single kind of organism at a time in a lab, commonly referred to as culturing. But even today, most microorganisms can’t be cultured. And, with the organisms scientists could culture in the early ’80s, they could only be identified or described using scraps of data, like the shapes of cells under a microscope.
In the 1970s, when the ability to sequence RNA and DNA was nascent and painstaking, microbiologist Carl Woese pioneered a method to sequence a short stretch of RNA that helps make a piece of the ribosome, the cellular protein factory. Unlike genes for feathers or photosynthesis, which are only found in some species, the ribosome is crucial to all known life forms. With ribosomal RNA (rRNA) sequences from cultured microorganisms, Woese could now definitively identify different species. He could also learn about how they were connected through evolution. Previous trees of life were based on observable characteristics, but Woese constructed a new tree based on the rRNA sequence, with three main branches: eukaryotes, the group that includes plants, fungi, and animals; bacteria; and a previously unknown group of single-celled organisms known as archaea.
In the 1980s, Pace had the crucial insight that you didn’t need to culture an organism to study it: You just needed to be able to get a small piece of RNA from the cell’s protein factory, rRNA, and sequence it.
Pace used water samples taken from a natural environment, going after the unculturable “extremophiles” from hot springs and hydrothermal vents—environments you can’t mimic in a lab. He isolated the mixture of all of the RNA from all the organisms in the sample and sequenced the short stretch of rRNA. The differences he saw in rRNA sequences within the same sample represented different lineages of microbes. The more similar the sequence, the closer together the organisms were on the tree of life. The more divergent the sequences, the longer since they had split off from an ancient common ancestor. Pace further refined Woese’s tree with his findings.
Banfield pioneered the next stage of studying microbes in the wild: genome-resolved metagenomics, an approach for putting together the complete genomes of organisms growing in natural environments. Banfield’s method starts with collecting samples that contain a mixed community of microbes. She then isolates DNA from the mixture. The isolated DNA is from potentially millions of cells comprising thousands of species. The DNA is broken into fragments short enough to sequence. Sophisticated computer algorithms then analyze the hundreds of millions of mixed-up DNA sequences, using the places where they overlap to build long, continuous sequences representing whole genomes.
When Banfield first suggested this approach, she was told it would never work. Imagine running a library’s worth of books through a paper shredder, then trying to piece them back together: The complexity would be too great. To reduce that complexity, she looked in Iron Mountain Mine, a Superfund site near Redding. The toxic drainage water from the mine is the most acidic water ever measured on Earth, and Banfield guessed that only a small number of organisms could live in that extreme environment. Now, instead of running a whole library through the shredder, it was just a few books. Still daunting, maybe doable.
She has not only identified new species but hung new branches on the tree of life, including organisms smaller than previously thought possible.
Banfield took a sample of a biofilm—a pink, slimy layer of mixed microorganisms—growing atop acidic drainage water. From the sample, she was able to put together nearly complete genome sequences for two organisms and partial genomes for three more. She introduced the technique and sequences in Nature in 2004. That paper had a profound influence on the field and is her most heavily cited work to date, with nearly 1,500 citations.
Looking at rRNA lets you differentiate between organisms and gives you an impression of how they are related in evolutionary time. Looking at a whole genome tells you not just what an organism is but what it can do: how it makes energy, reproduces, defends itself from attackers, what nutrients it gives and takes from a microbial community. By using genomic sequencing, Banfield and her group have found organisms whose genomes are so different than anything previously seen that they had gone entirely undetected by rRNA sequencing. The tree of life is larger—and more dominated by microbes—than we had ever known.
Since piecing together the first genomes from Iron Mountain, Banfield has put together complete genomes from bacteria, archaea, fungi, and viruses from microbial communities all over the globe, including Crystal Geyser in Utah, salt flats in Chile’s Atacama Desert, human saliva, dolphin saliva, hospital rooms, a bioreactor in South Africa, cow guts, freshwater lakes, oceans, soil, and material hundreds of meters down in the Earth’s crust.
Banfield and her team have found more than 1,500 new types of bacteria and archaea since pioneering genome-resolved metagenomics. She has not only identified new species but hung new branches on the tree of life, including organisms smaller than previously thought possible. These microorganisms are missing major genes needed for growing, making energy, and even reproduction. They likely survive by living as symbionts—or possibly parasites—in tightly knit microbial communities. Since traditional cultures consist of just a single type of microorganism, they automatically exclude these types of organisms. Her work has made it possible to find them.
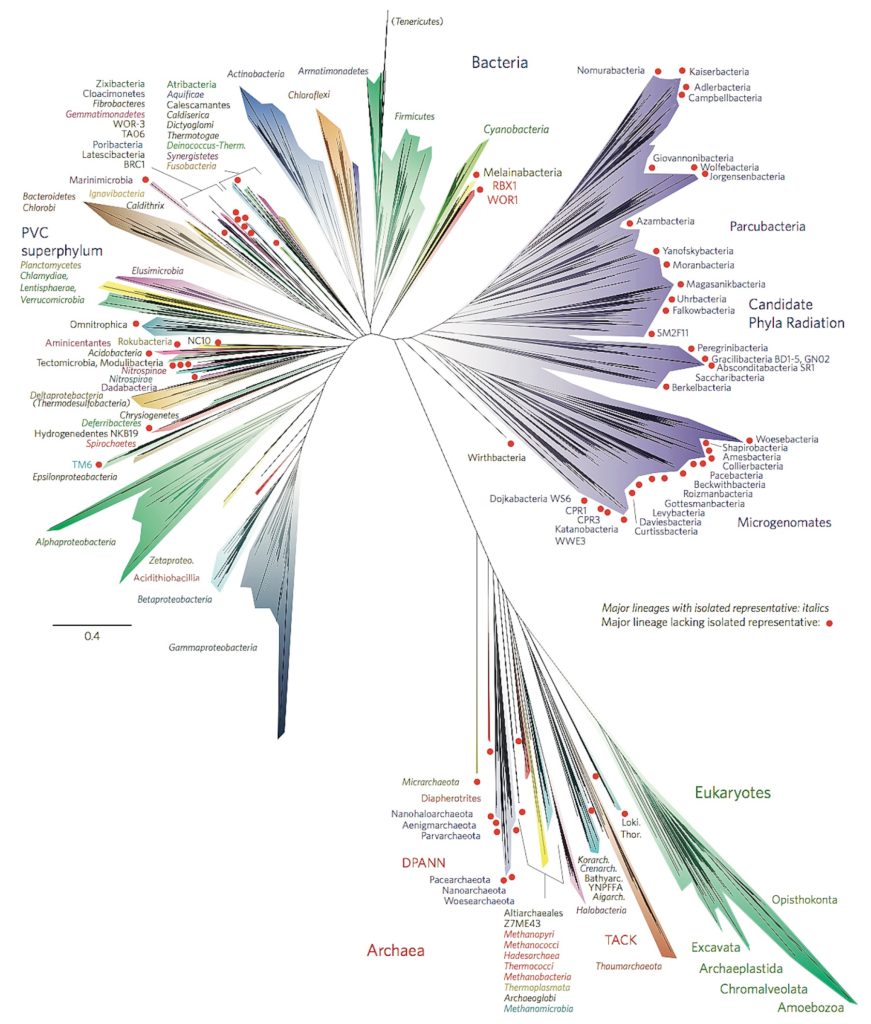
In 2016, Banfield and her team created a new tree of life. Instead of being based on one short stretch of rRNA sequence, this tree took into account sequences of 16 different proteins, yielding higher precision and resolution. Her analysis produced an evolutionary tree unlike any seen before: The branches are dominated by bacteria, especially nameless, unculturable bacteria. On her tree, bacteria represent more than two thirds of the genetic diversity on the planet. In other words, most kinds of living things are bacteria.
Beyond pioneering this revelatory technique, another key to Banfield’s ongoing discoveries is that she is hands-on with DNA sequence data. “When most people get a genome,” explains Alex Crits-Christoph, Ph.D. ’21, a recent graduate student in the Banfield Lab, “they look at it with a question they are trying to answer, or they quickly run a computer program to see what genes we already know about are there.”
The algorithms look for similarities, like a word search. But automation misses what is new, says Crits-Christoph. Banfield carefully scrolls through the rows of A’s, G’s, T’s, and C’s that make up genomic sequences, pulling patterns out of the seemingly opaque data.
As a Berkeley-trained Ph.D. geneticist, I can attest: I’ve never seen or heard of anyone doing this. It hadn’t occurred to me that you could just scroll through the thousands of letters that make a DNA sequence and see anything besides floaters. But Banfield can see things in the raw data.
“I think she has a good, I would call it kind of a sixth sense for interesting biological phenomena that you can observe at the level of genomics,” says Jennifer Doudna, Berkeley’s pioneering CRISPR researcher. “And I think a lot of top scientists have that type of ability. They just have a sense of interesting directions and questions to pursue.”
When Banfield looked at individual bacterial and archaeal DNA sequences, the palindromic repeats we now know as CRISPR would pop out. Before there was any understanding that CRISPR could be harnessed as a gene-editing tool, Banfield knew it was worth investigating.
“The reason I knew CRISPR was interesting, and pretty much was certain I knew what it was, is because we were working with reads,” says Banfield, referring to the raw DNA sequences.
She reached out to Doudna over email, and they met for the first time at Berkeley’s Free Speech Movement Café in 2006. “I remember that she brought a bunch of printed-out papers with her that talked about the sequencing data and what they might possibly be doing biologically. So it was pretty obvious that she was quite an amazing, creative thinker,” Doudna says.
Seeing a need for broader collaboration, Banfield convened a series of meetings on CRISPR, held in Berkeley between 2008 and 2012. As with the conference in Alta, she was able to convince world experts in disparate fields—chemists, microbiologists, even food scientists studying yogurt—to come together. Doudna would go on to win the 2020 Nobel Prize in Chemistry for her work on harnessing CRISPR for genome editing. Banfield had again identified something uniquely compelling that would change how we think of life itself.
This piece was not my first request to Banfield to write about her or her work, or the first profile she’s been asked to do. She is a private person, sometimes mistaken for shy at first, who tends to avoid being photographed when possible. She is uncomfortable with accolades, viewing her success as a result of group efforts. One lab member described learning of an award Banfield won only when she came to the lab with a film crew following.
It is true that Banfield does her extraordinary work in part by creating a network of students and postdocs who have gone on to shape the field of microbiology in their own right, collaborating with each other, some heading their own labs and biotech companies. “She puts people together as scientists, but as people, too. We work together because we’re better together—that’s Jill’s attitude,” says Kelly Wrighton, Ph.D. ’10, a former Banfield Lab postdoc, now heading her own lab in Colorado. “And we all sort of stick together. If anyone from the Banfield Lab contacted me, I would treat them as a brother or sister scientist. I don’t know if it’s deliberate, but Jill has created a lineage.”
“It’s very much deliberate,” Banfield says. “It’s a network like an old boys’ network, but for everyone.”
Banfield will always find time to look at data and write research papers with her lab members. Mentees describe her going through their manuscripts line by line, the figures graph by graph, even down to the colors used—teaching them to find the shape of the story hidden in the data and how to show it to the world.
Before the pandemic, Banfield brought current lab members to her off-grid property a couple hours northeast of Berkeley for annual retreats. They camped in the sprawling yard, woke up early to join Banfield on long, scrambling hikes, cooked and ate together, and talked about their lives outside of science. If they were lucky, Banfield would teach them how to make pottery, a longtime passion of hers that sometimes merges with her science.
In her office, there are sculptures of bacteria and their moon-lander-shaped viral predators, bacteriophages. When I interviewed her, she showed me a round, clay coronavirus she had sculpted, big as a volleyball. She pulled the top off to reveal an S-shaped strand of RNA inside.
Banfield says she sees a lot of similarity between understanding minerals and mineral structure and genomes. “Understanding the building blocks and how they’re put together was probably what really got me deeply enthralled about science.” With minerals, atoms are the building blocks. With genomes, it’s DNA.
At the heart of Banfield’s work ethic—the 12-hour days, the weeks-long fieldwork trips, the veritable hunger for data—is her drive to create. As an avid potter, she uncovers shapes she can divine inside blocks of clay. As a scientist, DNA is her raw material.
“She’s a creator first and foremost,” says Wrighton. “Data is the raw materials, right? Data is like that mound of clay. I think what Jill likes is working in that space and creating.… Seeing something in this amorphous space and doing actions and directions that pull something beautiful out of the heap.”
Hope Henderson holds a B.A. in biology from Brown University and a Ph.D. in molecular and cell biology from Berkeley. She works as a communications strategist at the Innovative Genomics Institute.